Functional parcellation of fMRI data.
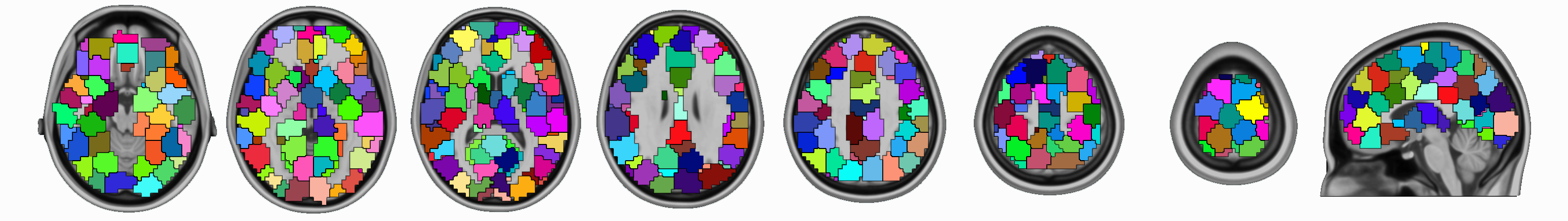
pyClusterROI is a set of python scripts for deriving a whole brain parcellation of functional magnetic resonance imaging (fMRI) data. The resulting regions are suitable for use as regions of interest (ROIs) in fMRI data analysis. This method was described and evaluated in the paper: Craddock, R. C.; James, G. A.; Holtzheimer, P. E.; Hu, X. P. & Mayberg, H. S. A whole brain fMRI atlas generated via spatially constrained spectral clustering, Human Brain Mapping, 2012, 33, 1914-1928 doi: 10.1002/hbm.21333.
This method employs a spatially-constrained normalized-cut spectral clustering algorithm to generate individual-level and group-level parcellations. The spatial constraint is imposed to ensure that the resulting ROIs are spatially coherent, i.e. the voxels in the resulting ROIs are connected. Using this package, clustering can be performed based on either the temporal correlation between voxel time courses, the spatial correlation between whole brain functional connectivity maps generated from each voxel time course, or a by spatial distance. Group level clustering can be achieved by either clustering the average of individual connectivity maps, or a 2-level approach in which single subject data is clustered, combined, and then submitted to another clustering. These methods require the specification of the number of clusters (ROIs) that the user would like to generate.
Based on the evaluations performed in the paper, 2-level group clustering using temporal correlation performed better than other methods. But the user might choose a different approach based on the specific analysis that is being performed. The group-mean approach requires much less computation, so it might be more appropriate for very large datasets. The user might prefer spatial correlation if they specifically want to optimize for the homogeneity of functional connectivity maps generated from within-ROI voxels. Evidence from the paper suggests that temporal correlation does a better job of optimizing the homogeneity of FC maps than does spatial correlation. Additionally, the number of clusters generated must be determined by the type of analysis to be performed. If the desire is to reduce the dimensionality to a low number while preserving functional homogeneity and interpretability, then a clustering in the range of 150 to 200 might be optimal. On the other hand, if the desire is to provide a modest amount of dimensionality reduction, but still preserve information present at the voxel scale, 600 - 1000 ROIs might be more appropriate.
Additional information can be found in a poster presented at the 2010 Annual Meeting of the International Society for Magnetic Resonance in Medicine.